The tetrapod body plan has withstood an enormous amount of variation on its basic theme, from fish to reptiles, amphibians and mammals. The limbs, too, vary enormously in their morphology, from fins to arms and legs, with two to as many as five digits. Molecular biology has provided enormous and growing insight into the genetic similarity onto which all this variation seems to have been superimposed, drawing a more and more detailed picture of the basic mechanisms of limb formation within tetrapods. As these basic mechanisms have been uncovered, they have lent much greater understanding to the evolution of limbs. It is this picture that I will outline in the following page.
The developmental mechanisms in the tetrapod limb are at once novel and extraordinarily conserved. Though it seems at first contradictory, portions of the development show apparent homology across most of the animal kingdom, while others seem the unique innovations of a common tetrapod ancestor. Any study of the evolution must attempt to make sense of this dichotomy. I will begin, then, with the novelties associated with the tetrapod limb, and then retreat to a more ancient evolutionary perspective to look at the apparent underlying homology.
The tetrapod limb can be divided into three parts from proximal to distal: the stylopod, zeugopod, and autopod. The first of these corresponds to the upper arm and leg of humans; the second to the forearm and lower leg, and the last to the hand and foot. The limbs are paired along the anterior-posterior axis, and always seem to develop at the same locations on the anterior-posterior axis of the body as defined by hox gene expression. For instance, the forelimb always develops at the most anterior region of Hoxc-6 expression in the trunk (Gilbert, 1997 and references therein). The bud then forms its own proximal-distal, anterior-posterior and dorsal-ventral axes. For more discussion of the development of these axes as a whole, click HERE. This page will focus on the development of these axes as they relate to evolution.
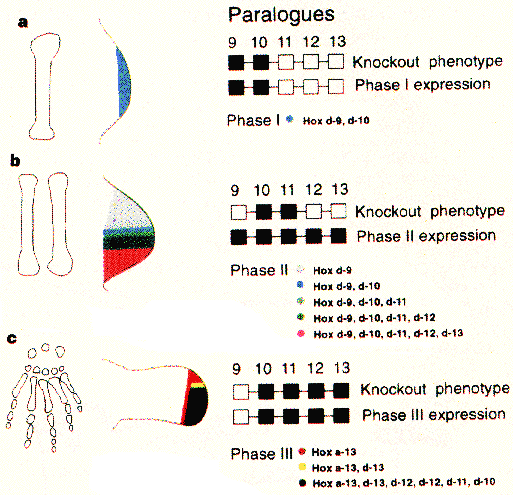
The three parts of the limb described above are not just anatomically apparent. They are developmentally important as well. Each of parts of the limb develops with a distinct phase of hox gene expression (Figure 1). In the first phase, for instance, hoxD-9 and hoxD-10 are coexpressed in the distal region of the limb bud (Figure 1a). This phase is important for proper stylopod devlopment (Shubin et al, 1997 and references therein). The second phase is important for the zeugopod, and involves genes 5' to those of the stylopod. Here hoxD-9 through 13 are expressed in a nested pattern which recapitulates the anterior-posterior polarization of the hox genes in the trunk. The most 3' gene, hoxD-9, is expressed over the entire limb bud, while more 5' genes' expression is increasingly restricted to the posterior of the limb, so that hoxD-13 has the smallest expression domain, restricted to the extreme posterior of the limb bud where it is coexpressed with the four other hox genes (Figure 1b). The third phase, in the presumptive autopod, reverses this pattern. Rather than the most 5' hox gene being the most restricted, as in the trunk and zeugopod, hoxD-13 has the broadest expression, while hox-9 through 12 expression is nested within its range. The phenotypes of hox gene knockouts correspond with this pattern as well (Figure 1c).

Figure 1. Hox gene expression in the developing tetrapod limb can be divided into three phases. Each successive phase involves prgressively more 5' hox genes, which assume more critical roles in development, as seen in the mutant phenotypes of murine knockouts for each gene (right). The first phase (a), involves hoxD-9 and hoxD-10 expression, and produces the stylopod. The second phase (b), which produces the zeugopod, involves all four hox genes -- hoxD9-13. The genes are expressed in a pattern reminiscent of hox gene expression in the trunk, in which expression of more 5' genes is nested inside the expression domains of more 3' genes. The third phase (c) reverses this pattern; it is the most 5' gene which has the most widespread expression. Figure from Shubin et al, 1997).
While the pattern of hox gene expression schematized in Figure 1 is useful in characterizing the peculiarities of autopod development, the in vivo expression appears somewhat different. While in phase 2 expression is neatly polarized along the anterior-posterior axis of the developing limb, in the autopod hox gene expression expands anteriorly in the distal portion of the bud, seemingly losing its anterior-posterior polarity (Figure 2). It seems this expression reflects a deflection in the axis itself (Figure 3b), so that from a molecular perspective, all the digits project posteriorly, even though morphologically they appear to extend on both sides of the axis. It is this deflection of the axis which seems to be a novel development within the tetrapods (Shubin, et al, 1997). Early vertebrates, for instance, do not show such a change. Instead, the "digits" in these fossils radiate to either side of an unwavering axis (Figure 3a). Phase 3 hox expression, in fact, is altogether absent from zebrafish unpaired appendages, and seems unique to tetrapods. Phase 2 expression, in contrast, in quite similar.
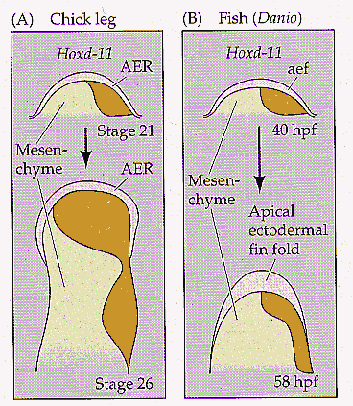
Figure 2. HoxD-11 expression expands anteriorly in tetrapods. In chick limb buds, hoxD-11 expression (orange) in phase 3 extends anteriorly nearly across the limb bud. In zebrafish unpaired fins, however, this expression is contained to the posterior of the limb bud. From Gilbert, 1997.
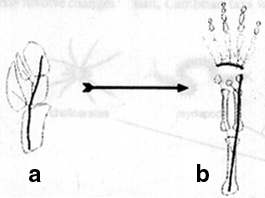
Figure 3. Evolution of the tetrapod limb correlates with a deflection of the limb anterior-posterior axis. In vertebrates, such as Panderichthyes, the anterior-posterior axis (black line) probably continued to the distal extreme of the fin, with digits extending anteriorly, or in other cases, to both sides of the axis. The autopod in tetrapods, in contrast, is characterized by an axis terminated before the distal end of the limb, with digits extending to the posterior side of this axis. From Shubin, et al, 1997.
A recent study looked at some of the elements necessary for regulating hox gene expression in the developing limb, attempting to tease out the details of how these phases are regulated, and how they may have evolved. Hérault and colleagues (1998) screened for conserved sequences surrounding the hoxD-12 gene, and found just two elements conserved between chickens, mice and zebrafish. Both elements are located at approximately the same locations in each species, and one, which the authors called RX, is conserved highly between all three species. The second, RXI, is more problematic. As shown in Figure 4, it is conserved at 56 percent over 206 bp between mouse and chick. Between chick and zebrafish, in contrast, the region of homology is much smaller, covering only 56 bp with 12 bp gap in the middle. Within this range the similarity is 60 percent. Even more perplexing, virtually no sequence similarity exists observed between zebrafish and mouse if it is calculated over the same region. The only similarity which does exist is in a 32bp subregion, in which the similarity is 53 percent. The implications of such similarity are not clear, but some possible interpretations are discussed below. First, however, it is helpful to consider the processes in which the authors implicated RXI.
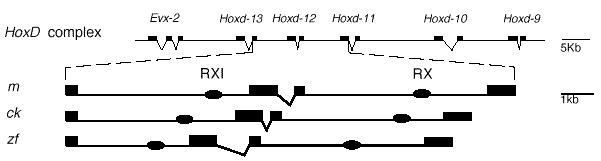

Figure 4. RX and RXI are conserved in zebrafish, mice and chickens. Both elements have been conserved in terms of location (top) and sequence (not shown for RX). However, the region of homology is much smaller between chick and zebrafish (56 bp) than between mouse and chick (206 bp). Between mouse and zebrafish the region of similarity is smaller still. (32 bp, in top line of sequence). From Hérault et al, 1998.
The authors deleted the RXI element by targeted mutagenesis.They found that mice homozygous for the deletion (RXIdel) showed identical hoxD-10, -11, and -13 levels to wild-type (not shown). HoxD-12 expression, however, was downregulated in the trunk and presumptive stylopod, and virtually abolished in the presumptive zeugopod (Figure 5). Expression in the digits was unaffected.
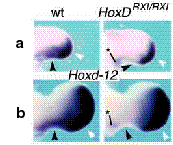
Figure 5. Deletion of RXI in mice results in abnormal hoxD-12 expression patterns. Both at 10.5 (a) and 11.5 (b) days post-conception, downregulation of hoxD-12 expression is apparent in the stylopod (*) and zeugopod (black arrowheads) of homozygous RXI-deleted mouse embryos. Expression in the autopod was unaffected (white arrowheads). From Hérault et al, 1998.
The changes in expression were complimented by analysis of mutant phenotypes in mice deleted for both RXI and nearby hox genes. Combining RXIdel with a null mutation for hoxA-11, for instance, resulted in deformities in distal portions of the radius and ulna, as well as the corresponding regions of the hindlimb. Furthermore, the same mice exhibited an increased number of conversions of the first sacral vertebra to lumbar. None of the combinations studied, however, including a triple inactivation of hoxD-11, -12 and -13, resulted in any increased phenotype in the digits, suggesting that this is the lone domain of hoxD-12 expression unaffected by the RXI region.
One would expect regulatory elements involved in autopod expression of hox genes not to be conserved in zebrafish. But RXI is involved in regulating hoxD-12 expression in all its domains except the autopod. Why, then, is the element so poorly conserved? A few possibilities exist. In isolation, the sequence data suggest that the chicken is more closely related to zebrafish and mouse than the latter two are to each other. This seems unlikely, since it contradicts the generally accepted view of verterbrate evolution as well as more specifically limb evolution as detailed above. An alternative hypothesis is that the only true homology in the three sequences exists between mouse and chick, and that the small amount of similarity with zebrafish is the result of convergent evolution. While it is true that small regions of homology might arise by chance in a large genome, the fact that this sequence is in approximately the same location in all three species makes this even more unlikely. A third hypothesis is that the zebrafish RXI element is in fact functionally equivalent to the other two, even though it has evolved differently. This hypothesis is supported by the fact that all of the base pairs conserved from mouse to zebrafish are identical to the sequence in chick as well. These nucleotides, then, are almost certainly of particular importance for all three species. Further, it suggests their function is most likely conserved as well. Thus, it seems this is the ancestral element, and that the tetrapod additions on it, while perhaps not relevant to digit formation, nonetheless represent a biochemical if not morphological novelty. Further, they are perhaps representative of the fairly minor changes in hox gene regulation and expression involved in the evolution of paired appendages. In order to be confident of this hypothesis, however, more research is necessary into the function of RXI in zebrafish.
The above data all emphasize, for the most part, the evolutionary novelty in tetrapod limb development. It has, after all, been a rather successful body plan, adapted to a variety of limb structures and functions, from hooves to fins to hands. But it seems inappropriate to discuss the novelty without also pointing out a deeper homology which unites the tetrapod limb to its ancestral limb, and it seems, to almost all known appendages.
Many of the genes involved in patterning the tetrapod limb bud have homologues active in patterning the corresponding parts of the Drosophilla version of the limb bud, the imaginal disc (Figure 5). Drososphila hedgehog, for instance, is expressed in the posterior portion of the imaginal disc, its homologue sonic hedgehog in the posterior of the limb bud. Sonic hedgehog induces BMP-2 expression along the anterior-posterior axis, which in turn is important in forming the anterior-posterior axis of the limb bud. In Drosophila, the BMP-2 homologue decapentaplegic serves the same purpose. Similarly, radical-fringe, serrate-2, wnt-7a, and Lmx-1 all have homologues fringe, serrate, wingless) or orthologs (apterous, a close relative of Lmx-1) involved in strikingly similar if not identical functions in patterning the Drosophjila limb (Shubin, et al, 1997 and references therein). These genes include signalling molecules and transcription factors several of which (e.g. wnt) seem to be easily adapted by evolution to a huge variety of situations. It seems extremely unlikely, because of this, that the two pathways evolved independently, even though it is not known how well this pathway is conserved in any of the phyla more recently diverged from either the arthropod or vertebrate lineage.
One gene which has been shown to have conserved expression patterns in many phyla is Drosophila distalless and its homologues, which are expressed at the distal end of developing insect and vertebrate limbs, ampullae and siphons of tunicates (urochordates), echinoderm tube-feet, annelid parapodia, and onychophoran lobopodia (Panganiban, et al, 1997).
The standard view of animal evolution holds that all these phyla evolved from a common ancestor resembling a flatworm (Figure 6). Since no known extant free-living flatworm has what one would consider an appendage, it is puzzling to figure out what the ancestral animal might have utilized these genes for. Several possibilities exist, however. The common ancestor may be farther removed evolutionarily and morphologically from extant platyhelmethes than the tree shown suggests. Thus it may have evolved a novel pathway for appendage-patterning which has been conserved since then in both the deuterostome (vertebrates, echinoderms, etc.) and protostome (arthropod, mollusc, etc.) lineages. Alternatively, there could be some inherent property of the way this pathway was utilized in the flatworm ancestor which made it easily adapted to appendage development. The pathway need not have been obviously related to any appendage per se, but merely possess some property in its regulation, position, or availability to evolutionary selection which led to its being independently co-opted at least twice to appendage development. According to this theory, most of the structure of the pathway was present in this common ancestor, though involved in a completely different process. A search for homologues of distalless and wnt in flatworms and even cnidarians would be of great help in supporting one of these hypotheses. Only with some insight into the early evolution of these genes can we come to a fuller understanding of the origin of the limb.
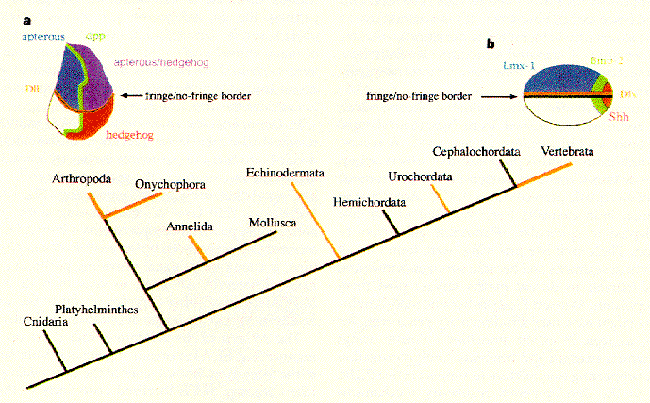
Figure 6. Various members of the appendage-patterning apparatus are conserved across the animal kingdom. Distalless homologues (orange, bottom) have been found in animal phyla thought to have diverged from common ancestors very early in animal evolution. Furthernore, a surprisingly large number of genes are expressed in similar areas of the developing limb buds of chicks (b) and the imaginal discs of Drosohpila, despite the apparent early divergence of these two lineages. These examples of the conservation in appendage devlopment suggest that virtually all animal appendages may all be formed through homologous mechanisms.
Panganiban, G., Irvine, S.M., Lowe, C., Roehl, H., Corley, L.S., Sherbon, B., Grenier, J.K., Fallon, J.F., Kimble, J., Walker, M., Wray, G.A., Swalla, B.J., Martindale, M.Q., and Carroll, S.B., 1997. The origin and evolution of animal appendages. Proc. Natl. Acad. Sci. USA 94(10):5162-6.
Hérault, Y., Beckers, J., Kondo, T., Fraudeau, N. and Duboule, D., 1998. Genetic analysis of a hoxd-12 regulatory element reveals global versus local modes of controls in the HoxD complex. Development 125: 1669-1677.
Gilbert, S. F., 1997. Developmental Biology. 5th ed. Sinauer Associates Inc., Sunderland. p.702-727.